Hi!
When you create a chain in the program, you are able to add the molecule name, type, sequence, and the number that the sequence begins at. Once you have created the numbering, is there a way to go back and change this information? For example, I create a protein to start at number 1 in the sequence. After a while, I need to fix the number so that the protein begins at number 20, not number 1. Just wondering if this edit is available in V3.
Thank you!
Domarin
Hi Domarin, We will add shortly a module to edit the chain. Meanwhile, do you need rename only one residue or `shift` the whole chain, with a new starting number?
To re-number the whole chain:
use chain.renumberResidues(offset, start, stop)
eg.
Assuming you have one chain in your project,
if your chain starts at 1 and finishes at 100 and you want restart from 20:
On the python console:
chain = project.chains[0]
chain.renumberResidues(20,1,100)
to rename only one residue:
r = project.getByPid(`MR:A.12.ALA`)
r.rename(`100`)
I hope it helps.
Luca
Hi Luca!
I just realized I never replied to this post. Yes, it did work! Thank you for the script.
Hi Luca!
I have another question about renaming residues. The macro works for renaming a chain. I also want to renumber my NMRchains. Is there a macro that will do this?
Thanks!
Domarin
Hi,
similar syntax, the command will be:
nmrChain.renumberNmrResidues(offSet, start, stop)
L
Hi Luca!
I am now trying to renumber the NMR chain in my project file using the command you gave `nmrChain.renumberNmrResidues(offSet, start, stop)`. I successfully was able to change the chain number using the first command you gave me. Using the nmr chain command, I get this error and am not able to figure out what is wrong. To give background, I have a protein that was originally numbered 1 to 290 and changed it to begin at 311 so my offset was 310. The following shows the command I used and the error I get:
chain = project.chains[0]
nmrChain.renumberNmrResidues(310,10,290)
---------------------------------------------------------------------------
AttributeError Traceback (most recent call last)
<ipython-input-30-1c430eed590f> in <module>
1 chain = project.chains[0]
----> 2 nmrChain.renumberNmrResidues(310,10,290)
<decorator-gen-216> in renumberNmrResidues(self, offset, start, stop)
/Applications/CcpNmrLauncher.app/Contents/Resources/ccpnmr3.0.2/src/python/ccpn/util/decorators.py in theDecorator(*args, **kwds)
326 self = args[0]
327
--> 328 application = self.project.application
329 blocking = application._echoBlocking
330 if blocking == 0:
/Applications/CcpNmrLauncher.app/Contents/Resources/ccpnmr3.0.2/src/python/ccpn/core/Project.py in application(self)
182 @property
183 def application(self):
--> 184 return self._appBase
185
186 def _initialiseProject(self):
AttributeError: `Project` object has no attribute `_appBase`
Hi,
my first thought, are you renumbering an nmrChain or a chain? In line 1 is chain = project.chains[0] . but in line 2 you call nmrChain. That nmrchain could be an old pointer to a previous session, or something else. Try replacing line one with nmrChain = project.nmrChains[0]
Hi Luca,
Thanks a lot, I was able to change the chain numbering but not the NmrChain. Which produces an unpleasant thing, I cannot change my assignment: e.g. A1 → A100 gives the error:
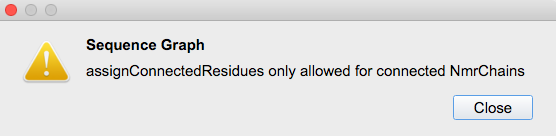
try the macro Renumber unconnected nmrChain
Hello,
I also have the same problem with nmrChain not connecting (with several gaps in between). I tried the latest macro, but it doesn’t seem to change anything. Is there another solution?
I would also like to know if it is possible to have a CS table in Sparky format in ccpn V3?
Thank you!
Regards.
Ons
Hi Ons,
we have now integrated the renumbering of Chains and NmrChains into the GUI. If you right-click on an NmrChain in the sidebar you can select Renumber NmrChain and it will do this and it won’t mind whether there are any gaps or not.
We don’t offer Sparky export. But if you open a Chemical shift list as a module, you can export it in Excel or .csv format and then you can reformat your columns if you like.
Vicky
Hi Vicky,
Thank you very much for the answer.
Indeed, I see the option but it asks only for the offset and doesn’t change anything if set it to any value … Any solution?
Thanks ! ![]()
Ons
Hi Ons,
that seems strange. If I set the offset to 5, then all the SequenceCodes for that NmrChain are increased by 5 when I click on Apply. You can also set the offset to a negative number, if you want to decrease the numbers.
Perhaps I am misunderstanding what you are trying to do?
Vicky
Hi Ons
However, we do offer NEF output and the latest released version of NEF-Pipelines which I code can translate NEF shift list to sparky ones after I rested the code from the attic. Steps as follows
- Please install NEF-Pipelines using the installer listed on its web page* using the script or uv [note I wouldn’t recommend using pipelines on windows… sorry]
[
- Test nef pipelines works, typing the command nef should give something like
nef
Usage: nef [OPTIONS] COMMAND [ARGS]…
Options:
–debug Enable debug output including stack traces
–install-completion [bash|zsh|fish|powershell|pwsh]
Install completion for the specified shell.
–show-completion [bash|zsh|fish|powershell|pwsh]
Show completion for the specified shell, to
copy it or customize the installation.
–help Show this message and exit.
Commands:
chains - carry out operations on chains [align]
csv - read [rdcs]
deep - read deep [peaks]
… [more lines removed for clarity!]
- Export a NEF file from analysisAssign
File → Export → NEF File
This should give you something like .nef, I am using the data from the Sec5Part4 CCPN project as an example. So it is in a file called Sec5Part4.nef by default
- You can now export the ccpn shift list to a sparky shift list
nef sparky export shifts --in Sec5Part4.nef --out - > Sec5Part4.txt
This should give you an output file that looks something like this
Group Atom Nuc Shift SDev Assignments
R3 CA 13C 55.551 0 1
R3 CB 13C 31.219 0 1
R3 H 1H 8.446 0.002 1
R3 N 15N 122.019 0.006 1
Q4 CA 13C 53.532 0.004 1
Q4 CB 13C 29.263 0 1
Q4 H 1H 8.379 0.002 1
Q4 N 15N 121.418 0.007 1
L7 CA 13C 53.996 0.015 1
L7 CB 13C 45.397 0 1
L7 H 1H 8.688 0.002 1
L7 N 15N 126.114 0.009 1
…
Which can be read by rl as normal in sparky or used elsewhere
Notes
- there is currently no support for multiple chains, if you have unassigned chains @xxx or @- these are treated like ordinary chains and each chain goes into a separate file
- Residues with offsets such as -1 e.g. @128-1 are ignored unless you use --include-negative-residues
regards
Gary
- note there is an install_nef_pipelines script in the ccpn distribution I currently wouldn’t recommend as it needs some revision [working on it!]
Dr Gary S Thompson NMR Facility Manager
CCPN CoI & Working Group Member
Wellcome Trust Biomolecular NMR Facility
School of Natural Sciences
University of Kent, Canterbury, Kent, England, CT2 7NZ
![]() :01227 82 7117
:01227 82 7117
![]() : g.s.thompson@kent.ac.uk
: g.s.thompson@kent.ac.uk
orchid: ORCID
Hello,
After assigning my protein, I would like to renumber the residues to fit the biological numbering, leading to my residue 5 to become residue 1, and my last residue 65 to become residue 61. I tried to run the macro with : nmrChain = get(‘NC:A’)
offset = -4
It gives me an error message saying it cannot renumber residue 65 to 61 because there is an existing assignment (and I am guessing it would do the same for the entire sequence). I tried also to just run in the python console :
chain = project.chains[0]
chain.renumberResidues(1,5,61)
For both I got the same essor message:
ValueError Traceback (most recent call last)
File ~/.ccpn/macros/macro_chain_renumber.py:25
23 for nmrResidue, newValue in zip(nmrResidues, newValues):
24 print(f’Renaming {nmrResidue} to → {newValue}')
—> 25 nmrResidue.rename(*newValue)
File ~/Documents/NMR/ccpnmr/miniconda/lib/python3.10/site-packages/decorator.py:232, in decorate..fun(*args, **kw)
230 if not kwsyntax:
231 args, kw = fix(args, kw, sig)
→ 232 return caller(func, *(extras + args), **kw)
File ~/Documents/NMR/ccpnmr/src/python/ccpn/core/lib/ContextManagers.py:1061, in renameObject..theDecorator(*args, **kwds)
1059 return _renameInner(application, args, func, kwds, self)
1060 else:
→ 1061 return _renameInner(application, args, func, kwds, self)
File ~/Documents/NMR/ccpnmr/src/python/ccpn/core/lib/ContextManagers.py:1069, in renameObject.._renameInner(application, args, func, kwds, self)
1066 with notificationBlanking(application=application):
1067 with undoStackBlocking(application=application) as addUndoItem:
1068 # call the wrapped rename function
→ 1069 result = func(*args, **kwds)
1071 if result is None:
1072 return False
File ~/Documents/NMR/ccpnmr/miniconda/lib/python3.10/site-packages/decorator.py:232, in decorate..fun(*args, **kw)
230 if not kwsyntax:
231 args, kw = fix(args, kw, sig)
→ 232 return caller(func, *(extras + args), **kw)
File ~/Documents/NMR/ccpnmr/src/python/ccpn/util/decorators.py:470, in logCommand..theDecorator(*args, **kwds)
468 # increase blocking
469 with notificationEchoBlocking(application=application):
→ 470 result = func(*args, **kwds)
472 return result
File ~/Documents/NMR/ccpnmr/src/python/ccpn/core/NmrResidue.py:1299, in NmrResidue.rename(self, sequenceCode, residueType)
1297 ll = self._project.getObjectsByPartialId(className=self.className, idStartsWith=partialId)
1298 if ll and ll != [self]:
→ 1299 raise ValueError(
1300 f’Cannot rename {self} to {self.nmrChain.id}.{sequenceCode}.{residueType or “”} - assignment already exists’)
1302 oldSequenceCode = apiResonanceGroup.sequenceCode
1303 oldResidueType = apiResonanceGroup.residueType
ValueError: Cannot rename NR:A.65.GLN to A.61.GLN - assignment already exists;
Anyone would have a quick solution for me ? I need to do it for 3 different chains (in different projects), so that the titration module can give me the CSP for the residues with the correct sequence number.
Best regards.
Laura
Hi,
I think the easiest way to get the behaviour you want should be via the GUI:
Expand NmrChains in the sidebar
Right click on the NmrChain you want to renumber
Select Renumber NmrChain
enter the desired offset and click apply.
I’ll look at why the other methods failed for you.
I’ve annotated the macro thread with an update warning of the limits to its use cases.
For the chain.renumberResidues() or nmrChain.renumberNmrResidues() functions, the issue is that you have misunderstood the meaning of the arguments. These are (offset, currentFirst, currentLast) and if you want to renumber a whole Chain or NmrChain you only need to give the offset and omit the other two.