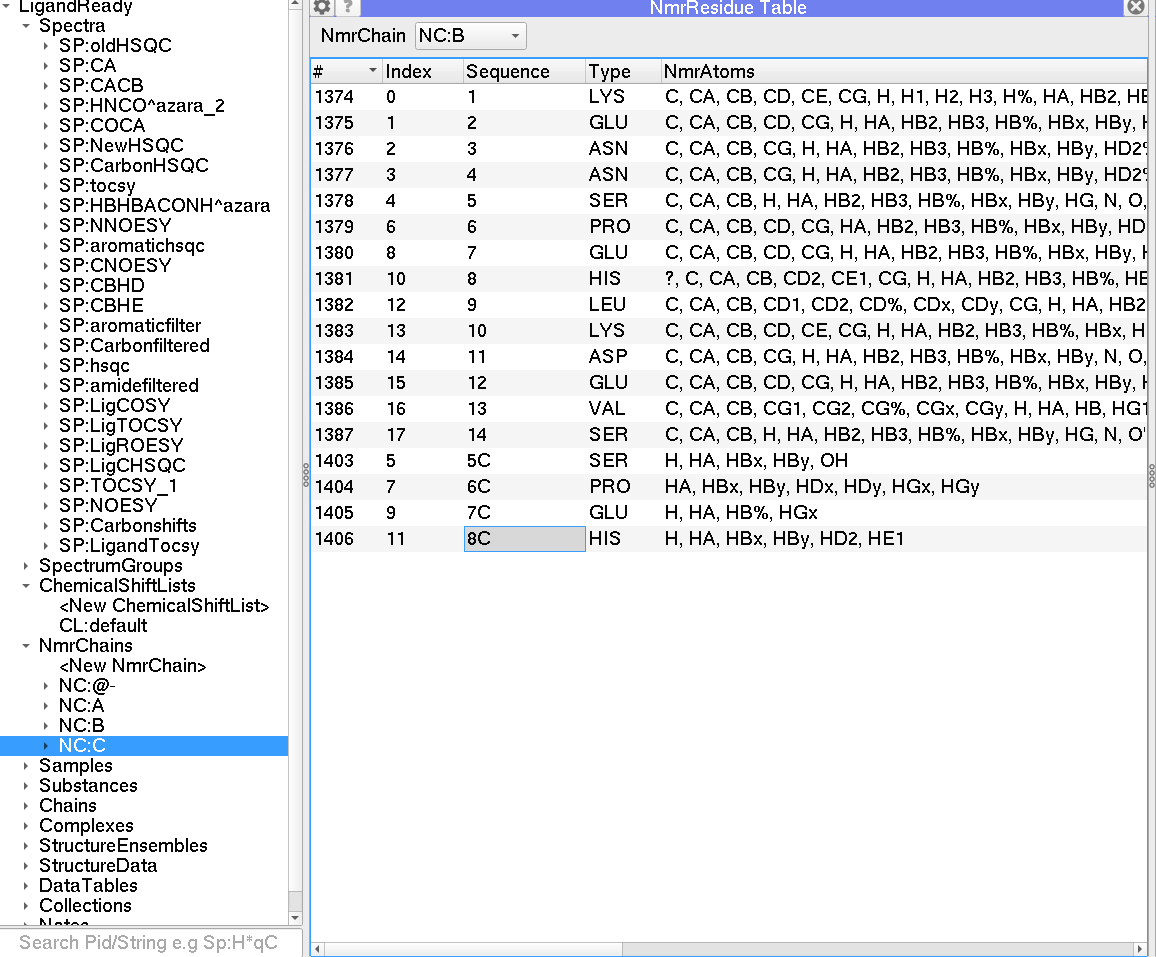
I have recently added a second ligand to my project called nmr chain “C”. C is identical to chain B and I want the chemical shifts Ive already assigned for B to copy over to C. I could clone the chain but the chemical shifts dont come with it. Is there any easier way to copy over the shifts from B to C. Ive searched everywhere but can find the option to do this.
Apologies if it’s straightforward.
My memory from doing this is I cloned the relevant peaks into a second peak list and then wrote a macro to change the chain for the peaks in those lists…
Dr Gary S Thompson NMR Facility Manager
CCPN CoI & Working Group Member
Wellcome Trust Biomolecular NMR Facility
School of Natural Sciences
University of Kent, Canterbury, Kent, England, CT2 7NZ
![]() :01227 82 7117
:01227 82 7117
![]() : g.s.thompson@kent.ac.uk
: g.s.thompson@kent.ac.uk
orchid: ORCID
Hi Joseph,
The answer will depend on what you mean to do e.g.
Do you want to assign all the peaks that are currently assigned to B to C also?
Do you want to populate just the ChemicalShifts of the NmrAtoms of C independent of any peak assignments (for example as a seed for generating predicted peaks)?
Something else?
Essentially I assigned my ligand and then subsequently realised it has two binding sites on my protein. So I needed to create another chain C with the same chemical shifts. So when I go back through the protein-ligand spectra with the peak assigner I have the option of clicking B or C chemical shifts to assign to it. But of course they will both be identical. I hope that makes more sense. So just copy the raw chemical shifts, dont change the assignments on spectra, I will do that manually.
So if there are two binding sites then in some cases technically there should be multiple chains if there is slow exchange. Basically you should have a chain for each distinct state. Most of the shifts except the binding sites and secondary effects will be the same. Its quite a hassle to maintain this sort of thing though so I would just maintain the shifts for the perturbed resonances in the two sets of peak lists for now and then add in the unchanged shifts if and when you need to deposit data..
Does that make sense?
regards
Gary
Dr Gary S Thompson NMR Facility Manager
CCPN CoI & Working Group Member
Wellcome Trust Biomolecular NMR Facility
School of Natural Sciences
University of Kent, Canterbury, Kent, England, CT2 7NZ
![]() :01227 82 7117
:01227 82 7117
![]() : g.s.thompson@kent.ac.uk
: g.s.thompson@kent.ac.uk
orchid: ORCID
This is the fix. Ill try and refresh my macro skills to chain the change for a designed peaklist ![]()