a very quick fix could be to select all peaks with cmd+A and hold down shift + the directional keys to move the peaks to a new position. Click on a spectrum display first to select it.
Zoom in-out to make more/less fine step adjustments while shifting
I lost a lot of peaks… my hncacb contains 950 pics, only 30 were imported, though all are well assigned in POKY…
I think this may be an issue to do with mapping the dimensions correctly and there may well be a bug which needs fixing. I remember looking into this a LONG time ago, but I’m not sure if we really fully resolved this.
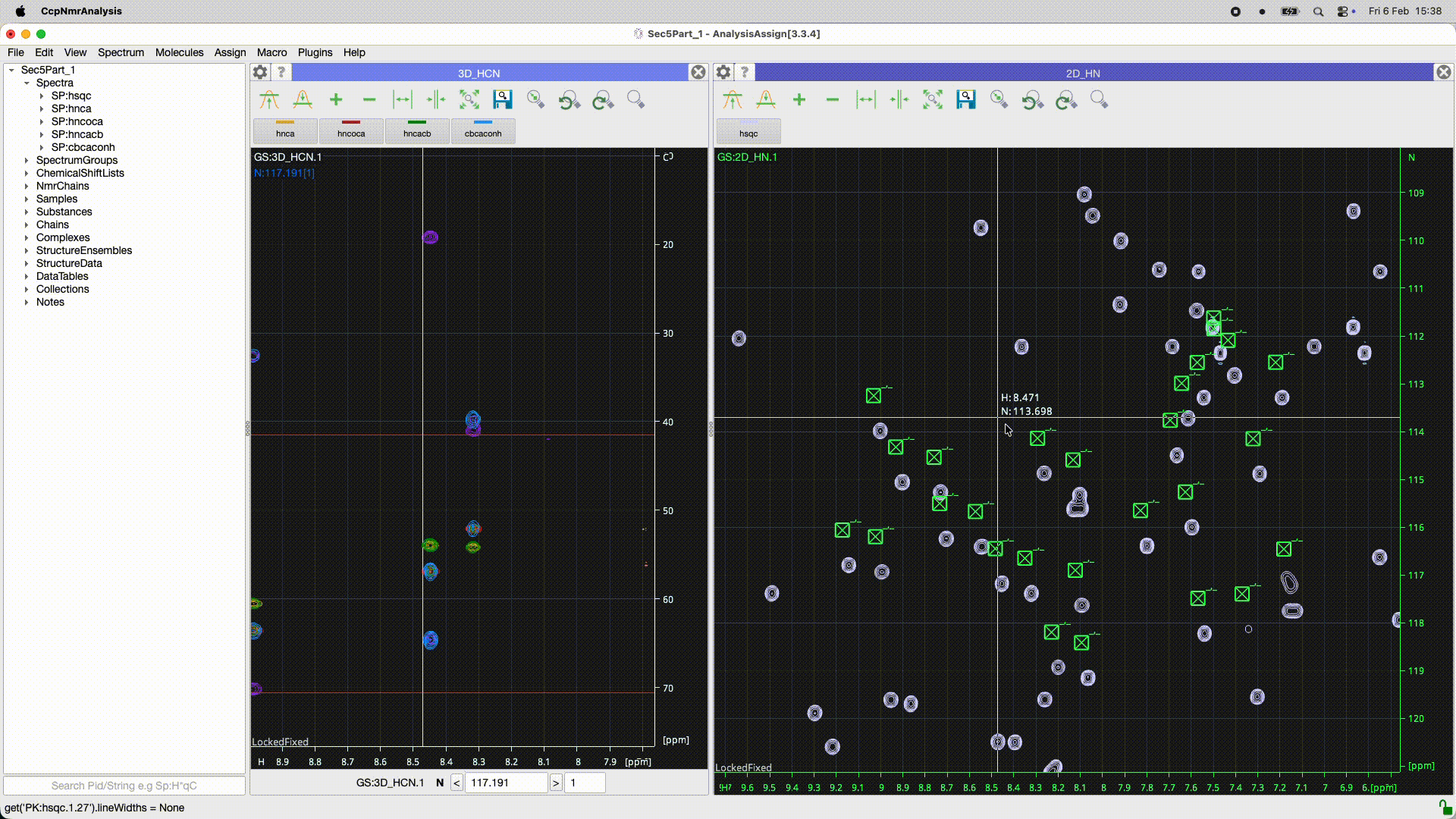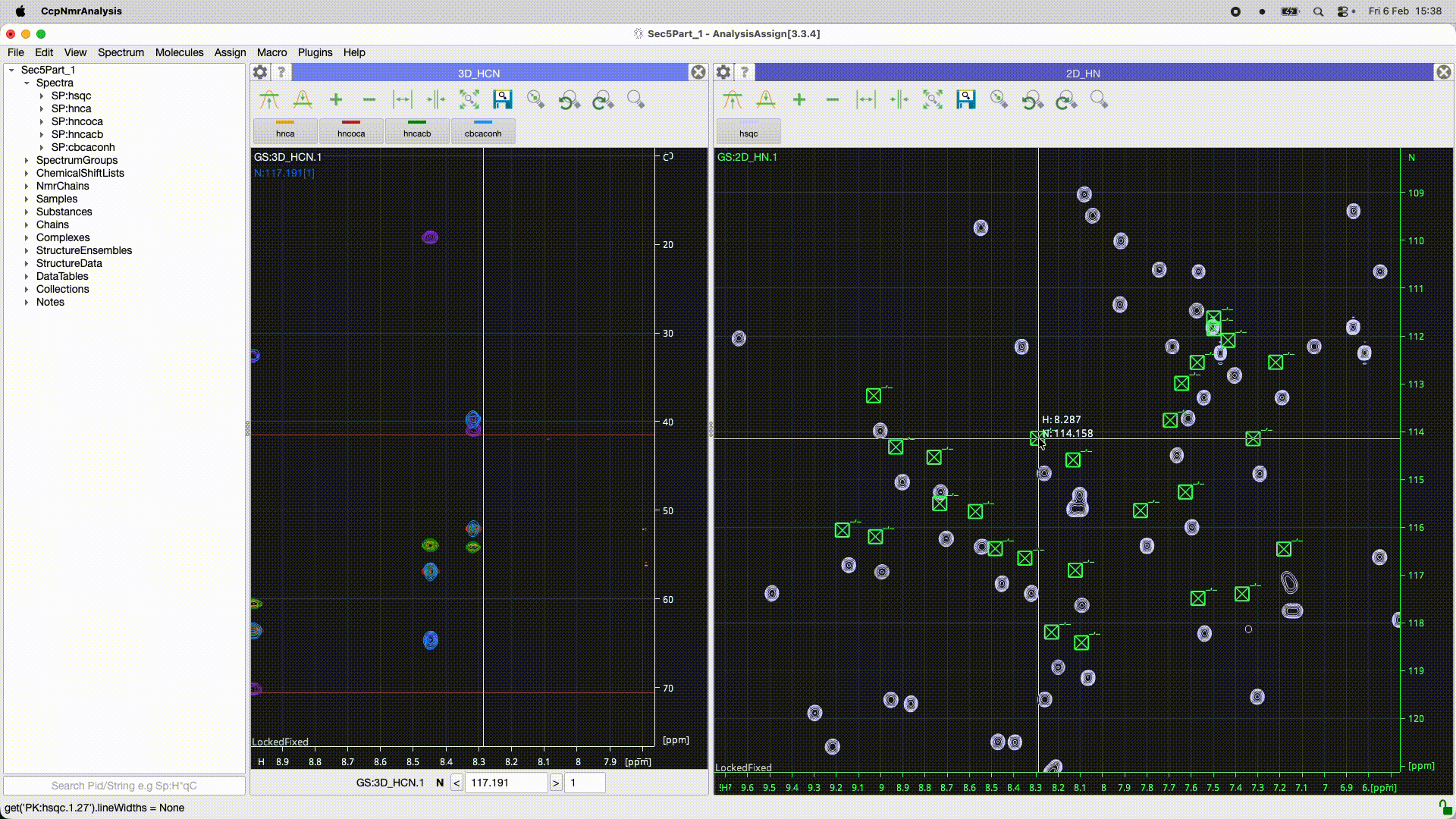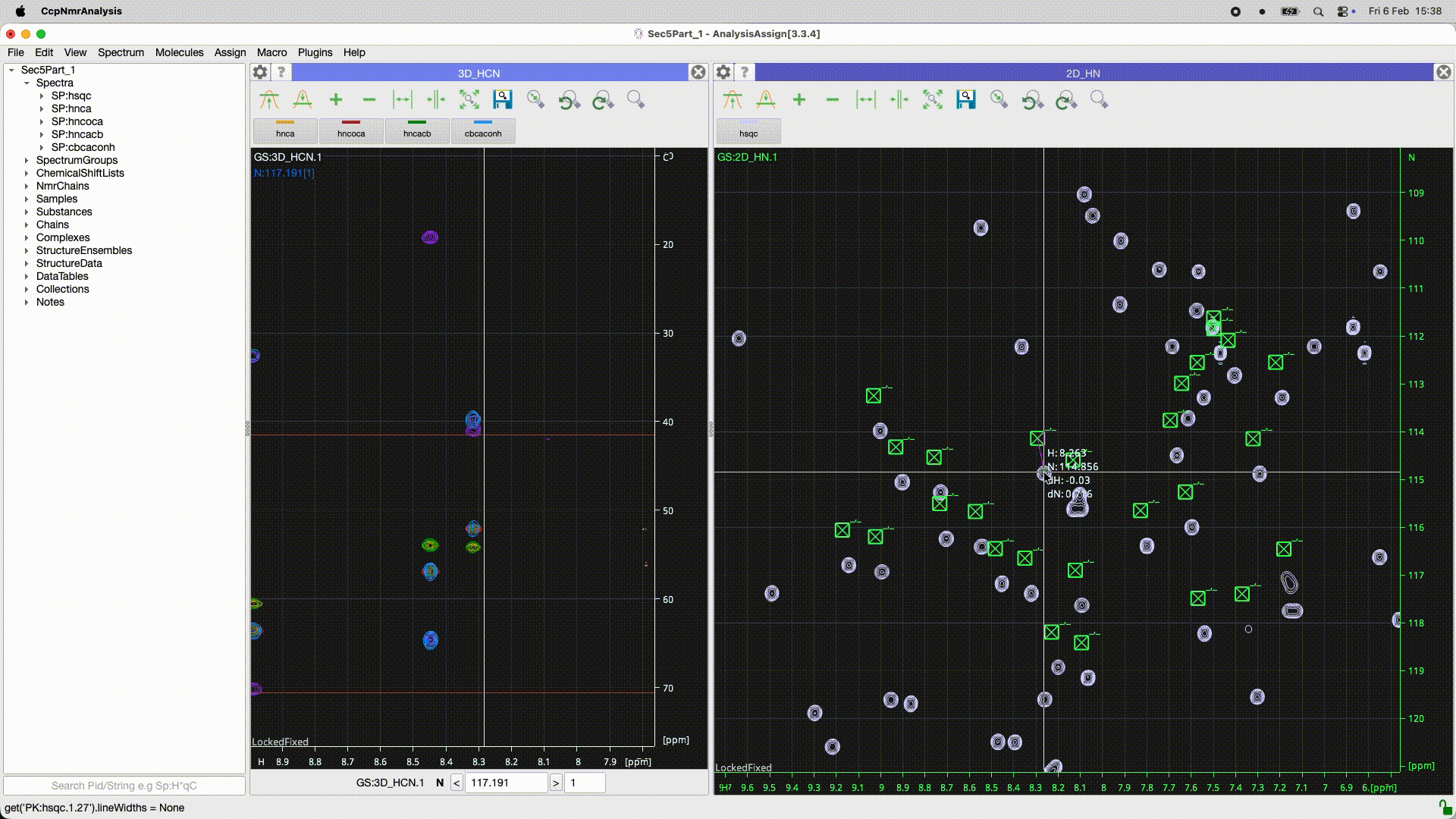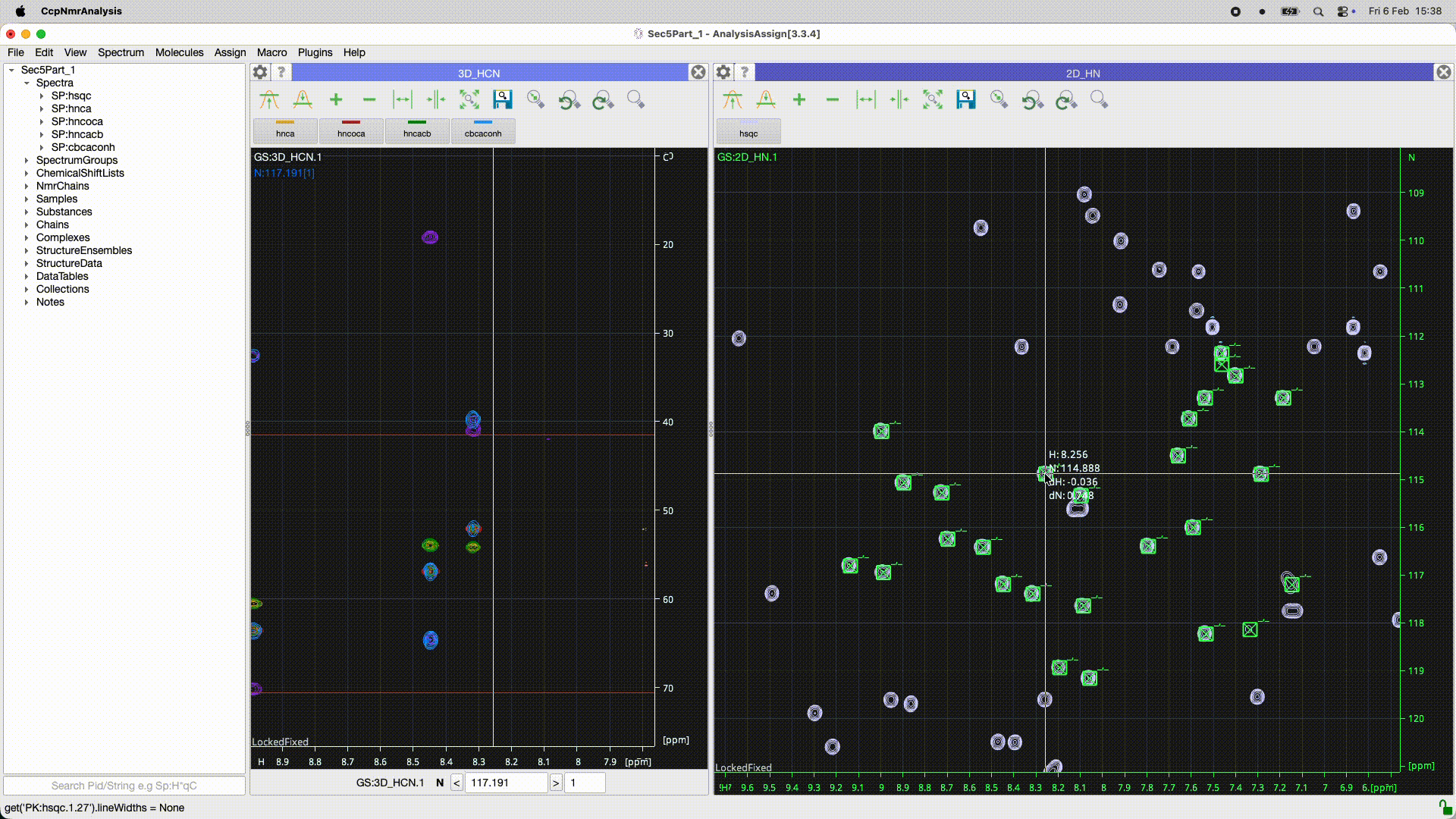
Can you work out what the program has done? In the 2D it looks as though there is simply a ppm offset for some reason. Is it the peaks or the spectrum which is offset? If you look at the peakList (nested under the spectrum in the sidebar) and open as module you should be able to see if the chemical shifts of the peaks are the same or not. You may need to re-reference the spectrum (had you done that in Poky?).
In the 3D, could it be that it has got the dimensions muddled up? Is there anything in peak 31 that would suggest why the program got stuck at that point? Again, looking at the Chemical Shifts of the peaks might enable you to see whether it has muddled the dimensions.
Vicky
Even better you can select all the peaks [control or apple A ] and do a right mouse drag on one of the peaks that shows the move peak line and if you move the line for that one peak to the correct position all the rest will go as well…
See the animated gif below if it will upload
regards
Gary
Dr Gary S Thompson NMR Facility Manager
CCPN CoI & Working Group Member
Wellcome Trust Biomolecular NMR Facility
School of Natural Sciences
University of Kent, Canterbury, Kent, England, CT2 7NZ
![]() :01227 82 7117
:01227 82 7117
![]() : g.s.thompson@kent.ac.uk
: g.s.thompson@kent.ac.uk
orchid: ORCID
I did it for the 2D. Thank you.
INFO : Importing Sparky molecular chains: MK2short_100ms_4struct4ccp (CcpnSparkyIo.importSparkyMolecule:1172)
INFO : Importing Sparky molecular chain: resonances (CcpnSparkyIo.importSparkyMolecule:1200)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
INFO : Importing Sparky molecular chain: resonances (CcpnSparkyIo.importSparkyMolecule:1200)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
INFO : Importing Sparky molecular chain: resonances (CcpnSparkyIo.importSparkyMolecule:1200)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
WARNING: Incorrect resonance. (CcpnSparkyIo.importSparkyMolecule:1226)
INFO : Importing Sparky molecular chain: resonances (CcpnSparkyIo.importSparkyMolecule:1200)
I have 4 different conditions for the same construct. I guess this latter refers to the 4 sets of resonances specified in the .proj file.
I try to pick up which resonances failed (warning) for each set of resonances.
It would be very helpful to print the incorrect resonances for the user to correct, or at least to inform him that these resonances cannot be imported.
INFO : Importing Sparky molecular chain: resonances (CcpnSparkyIo.importSparkyMolecule:1200)
INFO : Importing Sparky spectra: Ntrosy1_YFIVLMAT_MK2_50-364_170uM_pH62_25C_800D_240619 (CcpnSparkyIo.importSpectra:727)
INFO : Loading spectrum: /Users/ymonneau/Desktop/MK2/MK2_50-364/240619_YFIVLMAT_MK2_50-364_170uM_pH62_25C_800D/2/Ntrosy1_YFIVLMAT_MK2_50-364_170uM_pH62_25C_800D_240619.ucsf (CcpnSparkyIo.importSpectra:741)
WARNING: Axes error: axis 'H' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'N' is out of bounds (CcpnSparkyIo._testPermutation:836)
INFO : Importing Sparky spectra: Chmqc_YFIVLMAT_MK2_50-364_170uM_pH62_25C_800D_240619 (CcpnSparkyIo.importSpectra:727)
INFO : Loading spectrum: /Users/ymonneau/Desktop/MK2/MK2_50-364/240619_YFIVLMAT_MK2_50-364_170uM_pH62_25C_800D/3/Chmqc_YFIVLMAT_MK2_50-364_170uM_pH62_25C_800D_240619.ucsf (CcpnSparkyIo.importSpectra:741)
WARNING: Axes error: axis 'H' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'C' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Error importing peakList: Chmqc_YFIVLMAT_MK2_50-364_170uM_pH62_25C_800D_240619 0 (CcpnSparkyIo.importPeakLists:993)
INFO : Importing Sparky spectra: AROtrosy_YFIVLMAT_MK2_50-364_170uM_pH62_25C_800D_240619 (CcpnSparkyIo.importSpectra:727)
INFO : Loading spectrum: /Users/ymonneau/Desktop/MK2/MK2_50-364/240619_YFIVLMAT_MK2_50-364_170uM_pH62_25C_800D/5/AROtrosy_YFIVLMAT_MK2_50-364_170uM_pH62_25C_800D_240619.ucsf (CcpnSparkyIo.importSpectra:741)
WARNING: Axes error: axis 'H' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'C' is out of bounds (CcpnSparkyIo._testPermutation:836)
INFO : Importing Sparky spectra: HallCHarosm_YFIVLMAT_MK2_50-364_170uM_25C_800D_240619 (CcpnSparkyIo.importSpectra:727)
INFO : Loading spectrum: /Users/ymonneau/Desktop/MK2/MK2_50-364/240619_YFIVLMAT_MK2_50-364_170uM_pH62_25C_800D/7/HallCHarosm_YFIVLMAT_MK2_50-364_170uM_25C_800D_240619.ucsf (CcpnSparkyIo.importSpectra:741)
WARNING: Axes error: axis 'H' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'H' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'C' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'H_3' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'H' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'C' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'H' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'C' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'C' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Axes error: axis 'H_3' is out of bounds (CcpnSparkyIo._testPermutation:836)
WARNING: Error importing peakList: HallCHarosm_YFIVLMAT_MK2_50-364_170uM_25C_800D_240619 0 (CcpnSparkyIo.importPeakLists:993)
For the Chmqc, the “error importing peak list” resulted in:
3 missing peaks/assignment
1 peak unassigned
For the HallCHaro, it resulted in all 79 unassigned peaks not being imported
Would you be able to send us the .save file for some of the spectra/peak lists that didn’t import properly? Then we can hopefully work out what is going wrong and try and get this fixed.
Thanks,
Vicky
Hi Vicky,
Here is one extract from .save that failed after peak id 3 got loaded (as usual, linewidth/integral/fr lines stop the peak loading):
<ornament>
type peak
color 0 2 0 white
flags 0 0 0 0
id 1
pos 0.777 129.608 9.422
height 0.000 40637775872.000
type peak
color 0 2 0 white
flags 0 0 0 0
id 2
pos 1.792 121.073 8.958
height 0.000 97468530688.000
type peak
color 0 2 0 white
flags 0 0 0 0
id 3
pos 0.828 118.937 7.318
height 61966459449.939 68830945280.000
linewidth 86.888 79.512 48.314 fit
integral 9.5529e+12 ga
fr 0.026
type peak
color 0 2 0 white
flags 0 0 0 0
id 4
pos 1.074 118.918 7.317
height 59301689079.349 53851717632.000
linewidth 53.492 61.076 22.887 fit
integral 2.0480e+12 ga
fr 0.020
type peak
color 0 2 0 white
flags 0 0 0 0
Hi Yoan,
it looks like there were several of us making changes to the Sparky import routine and my last one may have got missed off and not put on the server. But Ed has just put it on now. With the latest updates, I am able to read in a .save file which has heights, linewidths and integrals. Though I notice that the integrals are currently not retained. I will have a look and see if it might be possible to change that.
Best wishes,
Vicky
Another important thing is the ability to extract peak note:
type peak
color 0 2 0 white
flags 0 0 0 0
note "Met2NH"
id 7
pos 119.003 7.320
height 0.000 110920925184.000
rs |L89|N| |L89|H|
[
type label
color 2 0 0 white
flags 0 0 0 1
mode 2
pos 119.003 7.320
label L89N-H
xy 119.052,7.112 7.191,118.708
]
User may have written important information about structure/assignment.