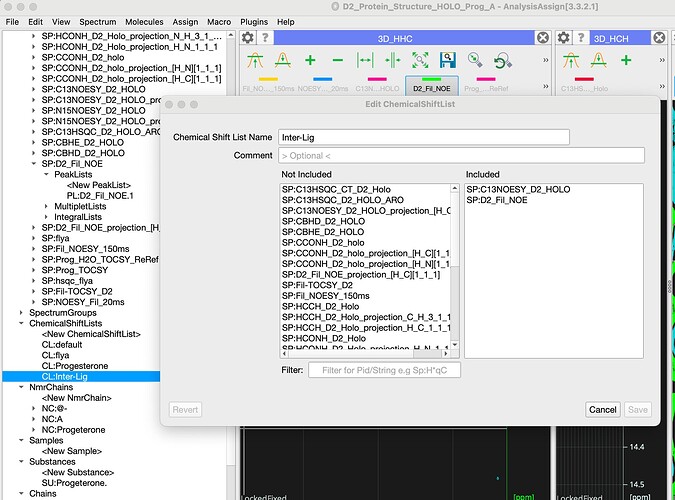
I am working on a ligand-protein complex. The ~117 residue protein binds a sterol. We have a model of the protein structure from an ARTINA/NMRtist run and I have imported the assigned 13C-/15N edited NOESY peaks into Analysis. I have also imported the ligand as a smile string to create a 2nd NmrChain B (see screenshot). I have a 3D w1-13C half-filtered, w2-13C-/15N-edietd NOESY and a w1-,w2-double 13C half-filtered NOESY (peaks are too broad in the TOCSY to be much use, due to exchange [?]). How do I set-up the project (peaklists) so that CCPN know that the signals in the 2D NOESY of the sterold bound to the protein come from the ligand resonances, and how do I tell CCPN that in the 3D w1 half-filted, w2-13C-edited NOESY that the signals in w1 are ligand resonances and the signals in w2 & w3 are protein resonances?
Thanks,
Mark